First cellular atlas of DNA-binding molecule could advance precision therapies
Biochemists at the University of Wisconsin–Madison have created the first atlas that maps where molecular tools that can switch genes on and off will bind to the human genome. It is a development they say could enable these tools to be targeted to specific parts of an individual’s genome for use in precision medicine, developing therapies and treating disease.
The study is published the week of Nov. 7 in the Proceedings of the National Academy of Sciences. The tools are polyamides, engineered DNA-binding molecules that are an important component of artificial transcription factors. Transcription factors — both natural and artificial — determine which genes are translated into proteins inside cells.
“We know that transcription factors bind to specific sites in the genome and when they misfire they drive many diseases, including cancers,” explains lead study author Graham Erwin, a former graduate student in the lab of Aseem Ansari, a professor in the Department of Biochemistry and the Genome Center of Wisconsin. “Using insights gleaned from this research, we hope to design polyamides that can bind to these same sites and outcompete the cancer-inducing factors, helping to repress that gene.”
Transcription factors work by binding to a particular gene and then recruiting the cellular machinery necessary to read it and manufacture the desired protein, or they can stop a protein from being created.
While polyamides had already been designed to attach to particular regions of the human genome and turn corresponding genes on or off, the new study answers lingering questions about where particular polyamides bind and ultimately function in the cell.
“Our big question was, where are these molecules going across the genome?” says Erwin, who is now a postdoctoral researcher at Stanford University. “With this study, we have a whole new understanding of how they read the genome.”
Using a technique called COSMIC (Crosslinking of Small Molecules to Isolate Chromatin), the researchers were able to create polyamides that would bind to DNA in human embryonic stem cells. Then, using a light-activated “handle,” the scientists could search for the location where the polyamide had bound to the genome. This enabled them to build an atlas of specific binding sites within the context of whole cells.
To their surprise, the researchers learned some polyamides can bind to DNA previously thought to be inaccessible.

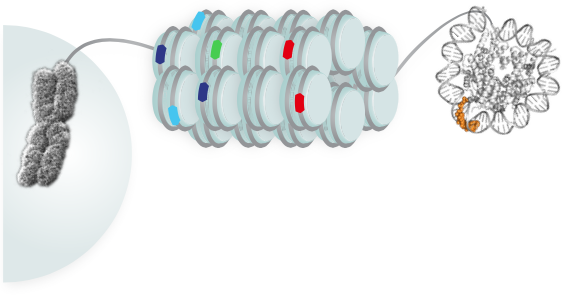
Each human genome, nearly five feet in length, must be highly packaged to fit inside the tiny volume of a cell’s nucleus. To accomplish this extraordinary task, cells wind up tightly most parts of the DNA that isn’t readily needed. At any given time, more than two-thirds of the DNA in the human stem cells the researchers used is packaged in this way.
The study team found some of the polyamides were bound to this off-limits DNA. While the DNA is unavailable to most transcription factors, polyamides may be small enough to reach it, the researchers believe.
“Being able to target a specific site in the genome is essential for the next generation of rationally designed therapies, and the lessons we’ve learned have changed the way we design molecules to target individual genomes,” says Ansari.
In a parallel study published Nov. 4 in the journal Angewandte Chemie, Ansari collaborated with Kan Xiong and Paul Blainey of MIT to visualize how these small molecules search long stretches of DNA for their binding sites. The study shows that these synthetic genome readers behave like “molecular sleds” and slide effortlessly across vast tracts of the genome.
Together the studies provide new insights into how these molecules locate their preferred target sites in the genome.
“For 15 years we’ve been working on this idea and now it seems we’re finally on the way to being able to intervene in a thoughtful way,” Ansari says.
Other authors on the PNAS “atlas” study include past or present members of the Ansari lab, members of the Department of Electrical and Computer Engineering, and researchers at the Morgridge Institute for Research.
This story was originally published on the UW-Madison News site.

